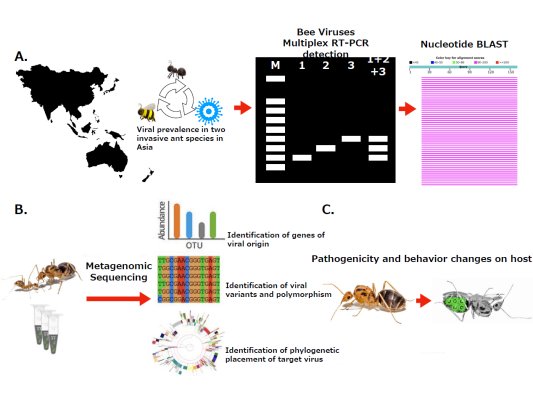
Ants often are infected by viral pathogens of honeybee origin possibly during their interactions with honeybee, suggesting that cross-species transmission of honeybee viruses may have occurred commonly in the ecosystem. While several studies have reported the presence of honeybee viruses in invasive ants, viral ecology and its potential effects on honeybee remain unclear and merit more research effort. To understand the dynamics of honeybee virus-invasive ant interactions, three major research themes were proposed as (1) to develop a specific and effective molecular method for detecting the presence of various known bee viruses in the two invasive ants in Asia; (2) to assess fundamental profiles of the honeybee viruses in the two invasive ant species including genetic diversity, infection patterns and geographic distribution; (3) to characterize changes at gene expression level, particularly those associated with behavior, through transcriptomic analyses in the invasive ants infected with honeybee viruses.
Here, samples from numerous geographic regions for two invasive ant species, namely yellow crazy ant (Anoplolepis gracilipes) and longhorn crazy ant (Paratrechina longicornis), were surveyed for the presence of six known honeybee viruses (including Acute bee paralysis virus, Black queen cell virus, Chronic bee paralysis virus, Deformed wing virus, Israeli acute paralysis virus, and Kashmir bee virus) using species-specific RT-PCR. The transcriptomic approach would facilitate our understanding of how invasive ants respond to honeybee viruses at gene expression level, shedding a new insight into virus-associated intrinsic responses of invasive ants to honeybee viruses. Results of this study represent the very first global survey effort and will serve as baseline information for evaluating the potential threat of the invasive ants as a source of “virus spillover” in honeybee and other pollinators that share close ecological associations with these ants.
ページ先頭へもどる
2019年4月18日作成