Invasive ants cause serious threats to naive community and biodiversity. In addition to chemical control, natural enemies, especially species-specific virus, may result in a devastating effect on their hosts, including immune interference and thus mortality. Recent results show that Solenopsis invicta virus-1 (SINV-1) infection reduces foraging intensity of fire ant colonies and changes their dietary preference, suggesting virus-ant interactions may play a role in shaping such behavioral responses. However, knowledge of virus infection in invasive ant species and the ant-virus interactions remains unclear, especially at physiological or molecular levels.
To understand the dynamics of virus-invasive ant interactions, I propose to develop an improved multiplex molecular method for simultaneously detecting seven known bee viruses in Argentine ants (Linepithema humile) and yellow crazy ant (Anoplolepis gracilipes) in Japan. The robust, fast and reliable screen methods (high-throughput metagenomics) are expected to help decode ecology of viruses in the two invasive ants. Moreover, the behavioral changes at gene expression level through transcriptomic analyses also will be characterized.
The project represents the first attempt to screen various bee viruses. The improved virus screening method would provide users critical information that help understand prevalence and diversity of the honeybee viruses in the invasive ants. The transcriptomic approach also is the first to demonstrate how invasive ants respond to honeybee-origin virus at gene expression level, shedding a new insight into virus-associated intrinsic responses of invasive ants.
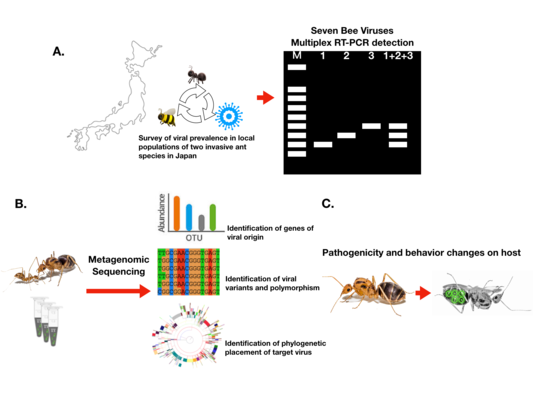 Fig. The schematic illustration of the study of virus-invasive ant interactions
Fig. The schematic illustration of the study of virus-invasive ant interactions
外来アリとウイルスの相互作用の模式図
ページ先頭へもどる
2018年10月29日作成


